Anne Paxton
April 2014—Talk to a few microbiology laboratories about why they feel the need to automate and you hear common themes: people, space, quality, and, most of all, time to detection. Microbiology may be late to join the bandwagon, but whether laboratories are making partial or full-scale moves to automate, they are dramatically making up for lost time, in all senses of the phrase. That’s because turnaround time savings are no longer measured in minutes. “Our goal is to be able to give some of these answers out in one to four hours rather than 24 hours, or much longer for some culture-based methods,” says Randall T. Hayden, MD, director of clinical and molecular microbiology at St. Jude Children’s Hospital in Memphis.

Horning
Instruments and modules on the market today have added new incentives for microbiology labs to automate at their own pace, says Ann Horning, MT(ASCP)SM, laboratory services manager for microbiology and point-of-care testing at Lancaster (Pa.) General Hospital. “If you look at microbiology two or three years ago, very few automation options were available.” In December, Lancaster General’s microbiology laboratory became the first in the U.S. to install the BD Kiestra InoqulA, delivering automated specimen processing and agar streaking via magnetic bead technology for liquid and non-liquid specimens.
Lancaster’s microbiology laboratory, which has 27 employees, does more than 15,000 tests per month, Horning says, noting that similar-sized area hospital labs in Reading and Harrisburg have also recently decided to automate specimen processing with the InoqulA or Copan Diagnostics WASP: Walk-Away Specimen Processor. “Microbiology labs are moving toward automation because of the focus on staffing and productivity. We can no longer afford to have medical technologists streaking plates.”
She finds that high-quality, standardized specimen inoculation is the biggest impact of the InoqulA. “How agar plates are streaked in the laboratory is variable with each technologist. Having a fixed amount inoculated onto each plate provides better quality outcomes. The staff has embraced this new technology and standardization.”
There has been one drawback, Horning says. “Because we are one of the first in the country to implement the InoqulA, the manuals and procedures that normally are available to us by the manufacturer are not yet available in the U.S. Specimen processing is performed in a biological safety cabinet in the U.S. to avoid infectious disease exposure to the employee. This technology has not yet been incorporated into the semiautomated component of the InoqulA. This is expected to be available by late summer.”
Once the biological safety cabinet is integrated into the InoqulA, the laboratory plans to move toward more around-the-clock plate reading. “Culture reading is performed now on day shift, but this technology will help us expand that to second and third shift.” By late summer, she hopes the laboratory will be performing culture reading 24/7 to improve turnaround time and patient outcomes.
At this point, the laboratory is not planning to reduce staff. “As we decrease staff time on manual plating by using the InoqulA, we have increased our molecular testing platform, so there has been some re-shifting of personnel,” she says.
Given the average age of medical technologists, the laboratory is expecting an outflux of personnel in the next five years, Horning says. “You’re going to want to have more support people to run InoqulA and do the less complex testing, while the medical technologists perform the complex duties. Most automated lines in the laboratory have aimed for staff reduction. This hasn’t yet happened in the microbiology lab, but we are probably heading in that direction.”
In fiscal year 2015, Horning says the laboratory hopes to expand its automation by acquiring the BD Kiestra Work Cell, a software and incubator solution that attaches to InoqulA to move plates in and out of special incubators that include digital imaging. “The one major thing it would impact is patient results in terms of turnaround time and length of stay, which has always been hard to measure. It will take some work to justify the substantial purchase, but hopefully the positive impact will allow us to purchase it.”
How close will that take the laboratory to total automation? “They have other pieces that can be attached, like the Bruker MALDI-TOF mass spectrometer. The Work Cell is a good addition for us as opposed to the TLA [total lab automation] because it will fit into our newly renovated laboratory without any further redesign.”
Meanwhile, some other U.S. laboratories are continuing their march toward total automation. At Evanston Hospital, part of NorthShore University Health System in the northern Chicago suburbs, the laboratory hopes to begin construction in April or May on renovations to prepare for its BD Kiestra TLA system, slated for delivery this summer, says Irene K. Dusich, MT(ASCP)SM, microbiology manager. Once the Illinois public health department has approved final drawings, “we’ll get installed in July, then probably take 90 days to validate it and fix the workflow the way we want it, so we’ll have it up by third quarter.”
With the Kiestra TLA, “the plates go directly from the line into the correct incubator, then they stay there and digital images are taken according to how we program it. Then, should we need to do a workup, the return line will bring the plate to whatever bench needs to work on it.”

Dusich
Dusich expects to see many things changing once the TLA system is in place, including a new ability for the laboratory to increase testing volume while reducing staff. “Microbiology is typically a day shift department, but a lot of work, especially outreach, comes in on the PM shift. Kiestra will alert us when plates are ready to be read according to how we have programmed the system. After an assessment period of at least six months, I can see us probably going to at least a two-shift operation for plate reading.” The laboratory now has 26 technical FTEs performing the work in microbiology. “We plan to reduce that by five FTEs with the Kiestra, and our hope is that the loss will be by attrition,” Dusich says.
Whether automation is the answer for all sizes of microbiology laboratories is not clear. At St. Jude Children’s in Memphis, for example, the microbiology lab would be considered small to moderate, with a staff of about 25, says Dr. Hayden. But in line with the hospital’s focus on catastrophic pediatric diseases, some of its tests are very high volume. “For example, we might do several hundred viral load tests for something like cytomegalovirus each month, which is comparable to what a very large academic center might do. On the other hand, since we don’t have an emergency room and only see pediatric patients, we do much fewer chlamydia or strep tests.”

Dr. Hayden
The lab automates individual tests when those tests become FDA cleared, but it does not use automation in the broad sense. “We have automated blood cultures, HIV is on an automated system, and we’ll probably be switching to automated testing for CMV viral load. But outside of molecular diagnostics we’re not a very highly automated lab, because for conventional bacteriology our throughput is pretty low,” Dr. Hayden says.
Automation has undeniable advantages in the setting of molecular testing. “If you adopt a sample-to-answer system, many of those will give an answer within an hour or maybe a little more. Some of these cartridge-based systems have limited to no sample prep required. So there may be a few minutes of hands-on tech time, and your turnaround time may drop dramatically. Because you’re not tied to batching, you may be able to give on-demand testing.”
Automation hasn’t reduced the number of staff needed in Dr. Hayden’s laboratory. “If we automate one assay it frees up part of some person’s time to maybe work on another assay. The margin of change in terms of reduced hands-on time isn’t typically enough to bump our staffing requirements downward, particularly given the ever-increasing need for additional tests.” Often labs that implement small-scale automated equipment, such as sample-to-answer PCR, have to pay a fair amount of attention to keeping throughput moving. “You can set up and walk away and it will run for an hour, but if you have four modules and 10 samples and they’re staggered, you’ll have to keep taking things in and out. Depending on how your space and workflow are organized, it might mean you need just as much hands-on time as running a batch system.”
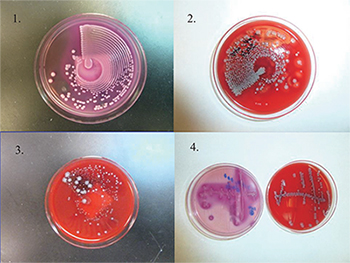
Culture plates (1–3) are streaked by using an automated system (Previ Isola). Fig. 1 is the MacConkey agar plate. Figs. 2 and 3 are the blood agar plate. Fig. 3 is a mixed culture with nice colony separation, which can be used directly for MALDI-TOF MS. For comparison, the culture plate 4 is streaked manually. Photos provided by Yun Wang, MD, PhD.
“On the bacteriology side,” Dr. Hayden says, “phenotypic testing involves relatively low-cost systems to begin with when you’re doing manual methodology. There are scaled systems for blood cultures and for automated bacterial ID and susceptibility testing and so forth, but some of the new technologies like MALDI-TOF are priced pretty high and may be harder for smaller labs to cost-justify.” However, because of the number of high-acuity patients at St. Jude, his lab was able to justify a MALDI-TOF and now has the system in validation.
For most labs, space continues to be scarce. “We’re always running out of space. If you are using modular systems and you need two or three or four modules, that takes up a bench position no matter what you do. Some manual methods aren’t ever going away, so basically you are adding on top of what you had before. We have an enormous box for our automated HIV, but we still need our individual cyclers to run LDTs. So I don’t think you can say we’re saving space. The hope is eventually for most systems we won’t need separate extraction, that front-end automated sample preparation will shrink, but at this point it’s just an admixture of systems. No one box does everything.”
To make some of the larger systems cost-effective and to justify taking them through the regulatory process, some manufacturers have targeted mostly high-throughput microbiology labs, Dr. Hayden says. “It’s just a function of how these things have evolved. But I think over time we’ll start to see more overlap and scalability, which will help us and help others.”
More automation is not always a good thing, he emphasizes. “Obviously, you need to ask not only the cost-benefit but also the impact of a new technology in your lab and the impact clinically. Will it work in your practice setting and for your patients and your technical staff? So it’s always a multifactorial decision.”
 CAP TODAY Pathology/Laboratory Medicine/Laboratory Management
CAP TODAY Pathology/Laboratory Medicine/Laboratory Management
